In a ``C-refinement'' the connection of a variable-pair is broken. The two
variables xi and xj are assumed to be independent and separated in each
subrelation. As an example assume the following model of the variable-space
![]() :
:
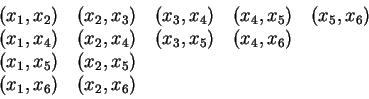
The final refined model after breaking the variable-pair (x4,x5) is:
The following is the complete C-refinement-lattice for an overall relation with
only 3 variables:
The second form of simplification is called ``G-Refinement''. In this more general refinement (it includes C-refinements indirectly by the use of several G-refinements) no direct links between variable pairs are broken. Instead one subsystem is replaced with all its sub-subsystems of complexity one less after removing redundant relations. In this way the combined effects of all variables in one relation are eliminated.
If, for example, only a specific combination of several variables triggers a medical condition, then this effect can only be represented in a relation containing all these variables together. After a G-refinement all variables still stay connected but not in the same relation and the model cannot represent the complex interaction of all variables.
As an example let us G-refine
![]() in our model:
in our model:
In theory we now have an algorithm (G-refinements) to enumerate all models and test them sequentially. In practise this is only possible for really small relations, as the number of models ``explodes'' with increasing number of variables [14, pg. 168]. For 3 variables there are only 9 possible models, but for 6 variables the number of models is estimated to be about 7 million.
This is the reason for using two different kinds of refinements. The more restrictive C-refinements can be used as "global search", while we look into the general G-refinements for local adjustments.
The procedure of global and local search is supported by several results that ``observed ... that reconstruction hypotheses have a tendency to naturally cluster into good and bad ones, i.e. into hypothesis with small and large distances'' [14, pg. 171]. Because of this ``natural clustering'' it is possible to search the enormous model space more effectively. We only need to look for ``interesting'' models in the neighborhood of already good models. One used strategy is to start with the original most complex model, refine it and test the resulting reconstruction hypothesis. From these models choose either the best, the best k, or the best p percent of the models for further refinement. Iterating this strategy down to the most simple model we get a pretty good idea about ``interesting'' models. If we only used C-refinements for the iteration we can use G-refinements to improve our ``local refinement search'' on these models.
Also mathematical optimization methods and Genetic Algorithms may be used for searching the model space. For these methods we either define a quality measure as a combination of complexity and accuracy or we only search a specific complexity level at a time and use the accuracy as quality measure (or fitness function). For Genetic Algorithms it also needs to be investigated how to encode models in character strings.